Advancing TAVI Outcome Prediction Through Privacy-Preserving Machine Learning
FLOTO leverages federated learning to develop robust predictive models for Transcatheter Aortic Valve Implantation outcomes while maintaining patient data privacy across multiple healthcare institutions.
Latest News
FLOTO @ DGK Mannheim
FLOTO will be on stage and with an interactive demonstration at the DGK annual meeting 2026.
April 2026
Website Update
We have updated the FLOTO website and given it a fresh new look!
February 2026
Ethics Ammendment
Handed in the ethics ammendment for extending the federation by several new institutions.
December 2025
World Heart Day
Contribution from the medical informatics initiative HIGHMed on the World Heart Day. View here.
September 2025
EHJ Digital Health
The FLOTO project and Sandy are featured in an article by the Europen Heart Journal Digital Health. View here.
May 2025
gesundhyte Article
We are featured in the gesundhyte.de magazine from 16th of December 2024, with a focus on transitional healthcare. View here.
December 2024
SYNERGIE Article
The DZG's SYNERGIE magazine has featured us in an article on Federated Learning that you can view here.
October 2024
EHEALTH Article
FLOTO is featured in the 05|2024 edition of the EHEALTHCOM magazine. View here.
May 2024
About the Project
Overview
FLOTO represents a groundbreaking approach to cardiovascular research, combining cutting-edge federated learning technology with clinical expertise in TAVI procedures. By enabling collaborative model training across institutions without sharing sensitive patient data, we aim to create more accurate and generalizable outcome prediction models.
Objectives
This project addresses the critical need for large-scale, diverse datasets in developing reliable clinical decision support tools while adhering to the highest standards of patient privacy and data protection.
Predictive Modeling
Develop accurate machine learning models to predict TAVI outcomes including mortality, complications, and quality of life improvements.
Privacy Preservation
Implement state-of-the-art federated learning techniques that enable multi-institutional collaboration without compromising patient privacy.
Clinical Integration
Create practical tools that can be seamlessly integrated into clinical workflows to support evidence-based decision making.
Knowledge Transfer
Establish frameworks for continuous learning and model improvement as new data and clinical insights become available.
Methodology
Our distributed learning framework allows hospitals to train models locally on their data, sharing only model updates rather than patient information. Standardized protocols ensure consistent data collection and preprocessing across all participating institutions, with rigorous multi-center validation to ensure our models generalize well across diverse patient populations and clinical settings.
Federated Learning Framework
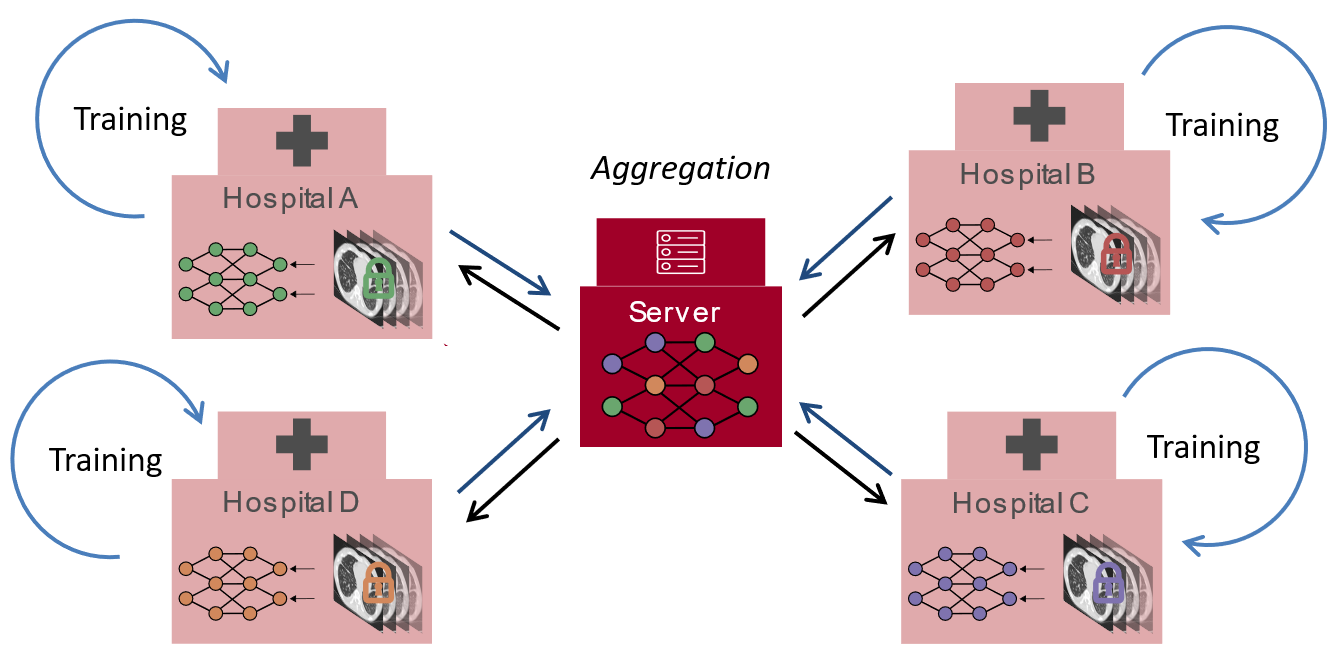
FLOTO on YouTube
Partner Institutions
FLOTO brings together leading cardiac centers, research institutions, and technology partners committed to advancing cardiovascular care through federated learning.
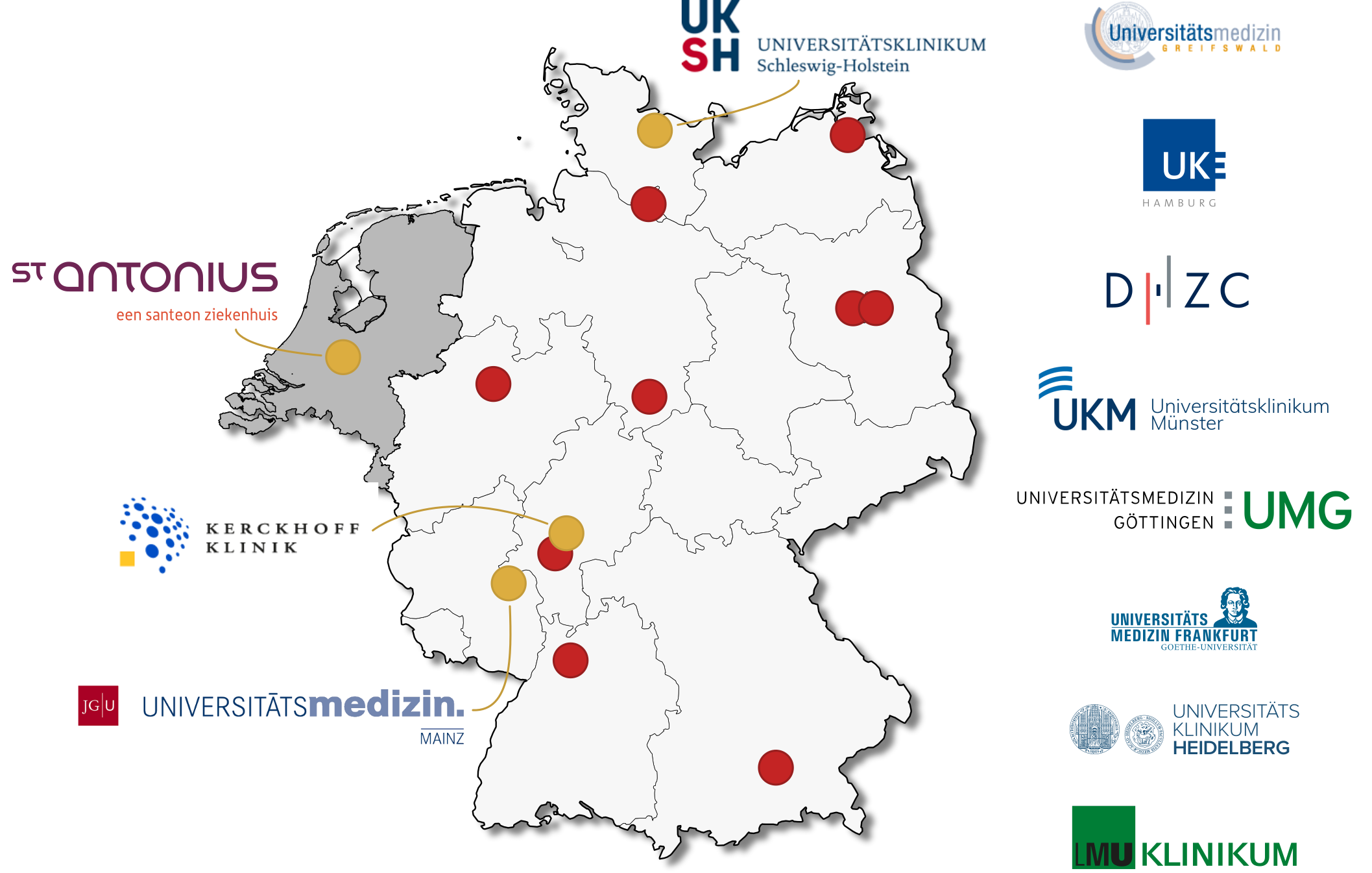
Principal Investigators & Research Team

Prof. Dr. Sandy Engelhardt
Department of Internal Medicine III: Clinic for Angiology, Pneumology and Cardiology, Heidelberg University Hospital, Heidelberg
Speaker

Dr.-Ing. Yannik Frisch
Department of Internal Medicine III: Clinic for Angiology, Pneumology and Cardiology, Heidelberg University Hospital, Heidelberg
Coordinator
M.Sc. Malte Tölle
Department of Internal Medicine III: Clinic for Angiology, Pneumology and Cardiology, Heidelberg University Hospital, Heidelberg
Dr. med. Florian André
Department of Internal Medicine III: Clinic for Angiology, Pneumology and Cardiology, Heidelberg University Hospital, Heidelberg
Prof. Dr. med. Peter Bannas
Department of Diagnostic and Interventional Radiology and Nuclear Medicine, University Medical Center Hamburg-Eppendorf, Hamburg
Prof. Dr. Norbert Frey
Department of Internal Medicine III: Clinic for Angiology, Pneumology and Cardiology, Heidelberg University Hospital, Heidelberg
Dr. rer. nat. Stefan Groß
Department of Internal Medicine B, University Medicine Greifswald, Greifswald
Prof. Dr.-Ing. Anja Hennemuth
Institute for Cardiovascular Computer-Assisted Medicine, Charité - University Medicine Berlin, Berlin
M.Sc. Nina Krüger
Institute for Cardiovascular Computer-Assisted Medicine, Charité - University Medicine Berlin, Berlin
Dr. Andreas Leha
Institute of Medical Statistics, University Medical Center Göttingen, Göttingen
Dr. Simon Martin
Institute for Experimental and Translational Cardiovascular Imaging, University Hospital Frankfurt, Frankfurt
Dr. med. Alexander Meyer
Department of Cardiothoracic and Vascular Surgery, German Heart Center Berlin, Berlin
Prof. Dr. Eike Nagel
Cardiovascular Imaging, University Hospital Frankfurt, Frankfurt am Main
Dr. med. Stefan Orwat
Department of Cardiology III: Congenital Heart Disease (ACHD) and Valvular Heart Disease, University Hospital Münster, Münster
Dr. med. Clemens Scherer
Cardiology, LMU University Hospital Munich, Munich
Prof. Dr. Stefan Simm
Institute of Bioinformatics, University Medicine Greifswald, Greifswald
Prof. Dr. Tim Friede
Institute of Medical Statistics, University Medical Center Göttingen, Göttingen
Univ.-Prof. Dr. Dr. med. Philipp Lurz
Department of Cardiology, University Medical Center Mainz, Mainz
Univ.-Prof. Dr. med. Philipp Wild
Department of Cardiology, University Medical Center Mainz, Mainz
Prof. Dr. med. Derk Frank
Department of Internal Medicine III, Cardiology and Internal Intensive Care, University Hospital Schleswig-Holstein, Kiel
Dr. med. Jakob Voran
Department of Internal Medicine III, Cardiology and Internal Intensive Care, University Hospital Schleswig-Holstein, Kiel
Dr. Martin Swaans
Cardiology Department, St. Antonius Hospital Nieuwegein, NL
Drs. Edgar Daeter
Cardiology Department, Heart Center, St. Antonius Hospital Nieuwegein, NL
Stan Benjamins
Cardiology Department, St. Antonius Hospital Nieuwegein, NL
Publications
Real world federated learning with a knowledge distilled transformer for cardiac CT imaging.
In: NPJ Digital Medicine
Abstract: Federated learning is a renowned technique for utilizing decentralized data while preserving privacy. However, real-world applications often face challenges like partially labeled datasets, where only a few locations have certain expert annotations, leaving large portions of unlabeled data unused. Leveraging these could enhance transformer architectures’ ability in regimes with small and diversely annotated sets. We conduct the largest federated cardiac CT analysis to date (n = 8,104) in a real-world setting across eight hospitals. Our two-step semi-supervised strategy distills knowledge from task-specific CNNs into a transformer. First, CNNs predict on unlabeled data per label type and then the transformer learns from these predictions with label-specific heads. This improves predictive accuracy and enables simultaneous learning of all partial labels across the federation, and outperforms UNet-based models in generalizability on downstream tasks. Code and model weights are made openly available for leveraging future cardiac CT analysis.
Multi-modal dataset creation for federated learning with DICOM-structured reports.
In: International Journal of Computer Assisted Radiology and Surgery (IJCARS)
Abstract: Purpose: Federated training is often challenging on heterogeneous datasets due to divergent data storage options, inconsistent naming schemes, varied annotation procedures, and disparities in label quality. This is particularly evident in the emerging multi-modal learning paradigms, where dataset harmonization including a uniform data representation and filtering options are of paramount importance. Methods: DICOM-structured reports enable the standardized linkage of arbitrary information beyond the imaging domain and can be used within Python deep learning pipelines with highdicom. Building on this, we developed an open platform for data integration with interactive filtering capabilities, thereby simplifying the process of creation of patient cohorts over several sites with consistent multi-modal data. Results: In this study, we extend our prior work by showing its applicability to more and divergent data types, as well as streamlining datasets for federated training within an established consortium of eight university hospitals in Germany. We prove its concurrent filtering ability by creating harmonized multi-modal datasets across all locations for predicting the outcome after minimally invasive heart valve replacement. The data include imaging and waveform data (i.e., computed tomography images, electrocardiography scans) as well as annotations (i.e., calcification segmentations, and pointsets), and metadata (i.e., prostheses and pacemaker dependency). Conclusion: Structured reports bridge the traditional gap between imaging systems and information systems. Utilizing the inherent DICOM reference system arbitrary data types can be queried concurrently to create meaningful cohorts for multi-centric data analysis. The graphical interface as well as example structured report templates are available at https://github.com/Cardio-AI/fl-multi-modal-dataset-creation.
Towards unified multi-modal dataset creation for deep learning utilizing structured reports.
In: Bildverarbeitung für die Medizin (BVM) 2024
Abstract: The unification of electronic health records promises interoperability of medical data. Divergent data storage options, inconsistent naming schemes, varied annotation procedures, and disparities in label quality, among other factors, pose significant challenges to the integration of expansive datasets especially across instiutions. This is particularly evident in the emerging multi-modal learning paradigms where dataset harmonization is of paramount importance. Leveraging the DICOM standard, we designed a data integration and filter tool that streamlines the creation of multi-modal datasets. This ensures that datasets from various locations consistently maintain a uniform structure. We enable the concurrent filtering of DICOM data (ie images and waveforms) and corresponding annotations (ie segmentations and structured reports) in a graphical user interface. The graphical interface as well as example structured report templates is openly available at https://github.com/Cardio-AI/fl-multi-modal-dataset-creation.
Content-aware differential privacy with conditional invertible neural networks.
In: Distributed, Collaborative, and Federated Learning, and Affordable AI and Healthcare for Resource Diverse Global Health (DeCaF FAIR) 2022
Abstract: Differential privacy (DP) has arisen as the gold standard in protecting an individual’s privacy in datasets by adding calibrated noise to each data sample. While the application to categorical data is straightforward, its usability in the context of images has been limited. Contrary to categorical data the meaning of an image is inherent in the spatial correlation of neighboring pixels making the simple application of noise infeasible. Invertible Neural Networks (INN) have shown excellent generative performance while still providing the ability to quantify the exact likelihood. Their principle is based on transforming a complicated distribution into a simple one e.g. an image into a spherical Gaussian. We hypothesize that adding noise to the latent space of an INN can enable differentially private image modification. Manipulation of the latent space leads to a modified image while preserving important details. Further, by conditioning the INN on meta-data provided with the dataset we aim at leaving dimensions important for downstream tasks like classification untouched while altering other parts that potentially contain identifying information. We term our method content-aware differential privacy (CADP). We conduct experiments on publicly available benchmarking datasets as well as dedicated medical ones. In addition, we show the generalizability of our method to categorical data. The source code is publicly available at https://github.com/Cardio-AI/CADP.
Presentations 2026
Sandy Engelhardt, UKHD: Image Synthesis, Federated Learning, and Vision-LLMs in Cardiovascular Medicine: Emerging Paradigms for Precision Care
@ Invited Guest Talk at AISCM Innsbruck
Sandy Engelhardt, UKHD: AI in structural heart today and future: Advancing TAVI outcome prediction through privacy-preserving machine learning
@ ABBOTT EDUCATION NETWORK - LIVE WEBINARS 2026
Tim Friede, University Göttingen: KI in der Gesundheitsversorgung und klinischen Forschung
@ Parlamentarischer Abend der LandesHochschulKonferenz Niedersachsen 2026
Presentations 2025
Sandy Engelhardt, UKHD: Multimodal federated learning for TAVI planning
@ ESC Digital and AI Summit 2025
Tim Friede, University Göttingen: Applications of AI/ML in clinical medicine
@ ASA Biopharmaceutical Section Regulatory-Industry Statistics Workshop 2025
Sandy Engelhardt, UKHD: Multimodal AI meets Engineering: Precision Medicine for Interventional Planning and Surgical Support
@ MICCAI CLIP Workshop Keynote 2025
Malte Tölle, UKHD: Multi-Modal Federated Learning from Heterogeneous Cardiovascular Data
@ ODELIA Summer School 2025
Sandy Engelhardt, UKHD: Multimodal Federated Learning & Image Synthesis for Cardiovascular Medicine
@ AI in Medicine by Fondazione Menarini
Sandy Engelhardt, UKHD: How to get AI bedside of aortic stenosis: AI-enhanced support to manage post procedural complications
@ ESC Main Congress 2025
Sandy Engelhardt, UKHD: Verteiltes Lernen von KI-Ansätzen für die Vorhersage von Schrittmacheabhängigkeit nach TAVI
@ DGK Annual Congress 2025
Contact Us
Project Coordination
For general inquiries regarding the FLOTO consortium
and partnership opportunities.
Technical Inquiries
For questions about our Federated Learning implementation
or data protocols.
Our Location
University Hospital Heidelberg
Im Neuenheimer Feld 420
69120 Heidelberg, Germany
